About me
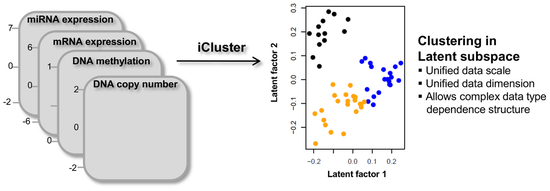
I am a thoughtful genomics data scientist statistician with strong programming skills in R and Python, and a formal training in both Computational Biology and Biostatistics. Currently, I work as a Principal Scientist at Incyte deploying ML models to mine for novel drug targets and making my work more accessible via data visualization and Shiny and FlexDashboard apps. Previously, I worked at Memorial Sloan Kettering Cancer Center and focused on methodological work in the field of Cancer Genomics.
The right hemisphere of my brain is into ceramics, painting, DIY crafts and biking. I am a minimalist and on a personal mission to reduce what goes in my trashbin. I also co-host a podcast on Computational Biology called Computationally Yours!
Education
MS Biostatistics, 2017
Columbia University
MS Computational Biology, 2010
Carnegie Mellon University
B.Tech Biotechnology, 2008
Amity University